Dispersal.jl is a collection of rules for organism growth and dispersal that can be used for modelling a wide range of ecological problems, and easily extended and combined with custom rules and other packages. Dispersal.jl extends DynamicGrids.jl, a high-performance grid-based modelling framework with clean syntax.
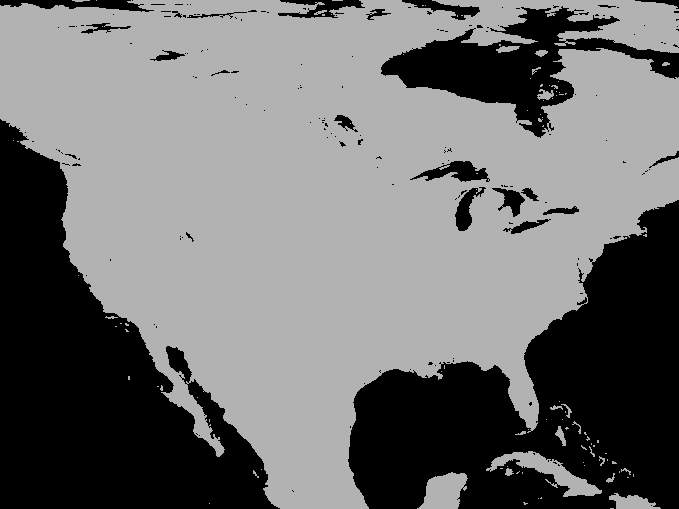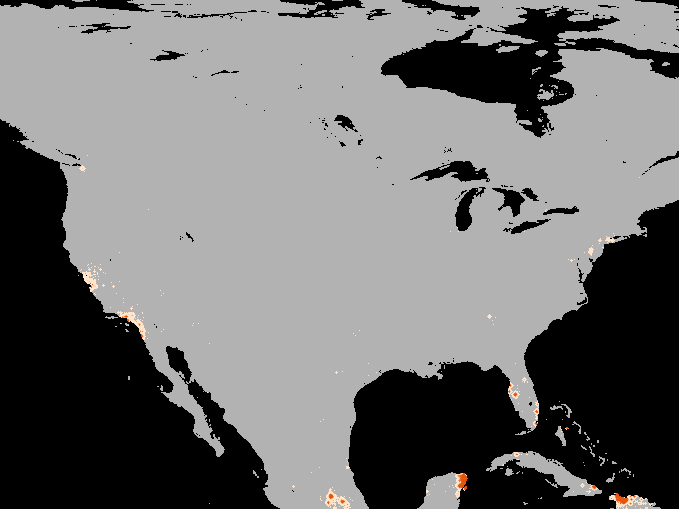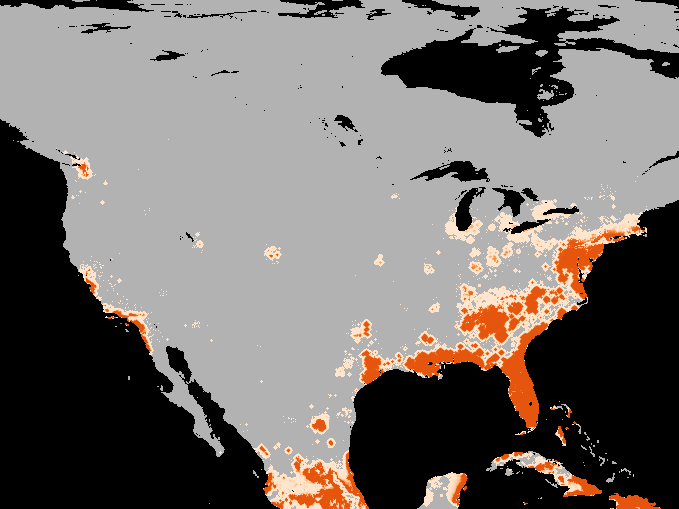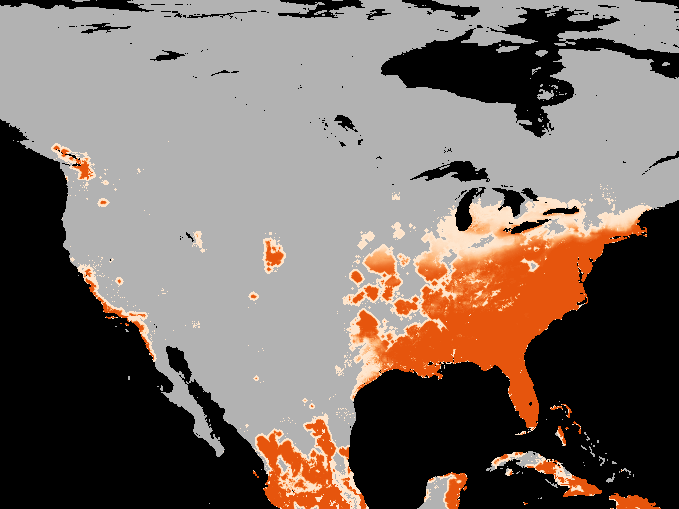
A simulation of the spotted-wing drosophola invasion of the continental United States as detailed in Maino, Schouten, and Umina (2021)
Growth rates, dispersal kernels, Allee effects, and randomised jump and human assisted dispersal rules are provided. These components can be combined into complex dispersal models. Custom rules can easily added and combined with the provided set. See the documentation for examples and the lists of included rules.
DynamicGridsInteract provides an interactive interface for atom and jupyter notbooks (InteractOuput), desktop (ElectronOutput) and online web applications (ServerOuput), where complete models, including your custom rules, can be manipulated during live simulations.
DynamicGridsGtk provides GtkOutput for a simple graphical viewer.
GrowthMaps.jl can efficiently generate summarised raster data for vital rates (e.g. intrinsic growth rates) based on higher resolution and shorter interval environmental data.